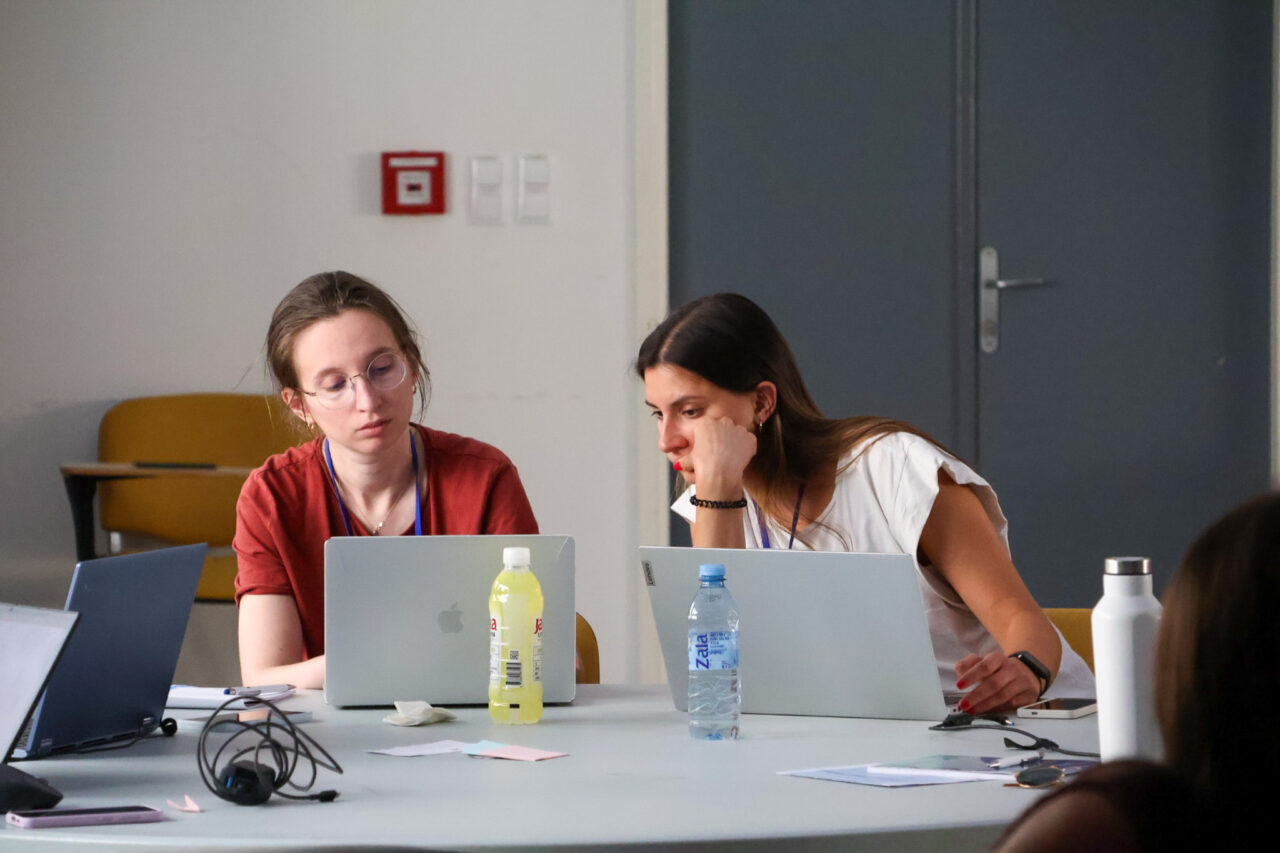
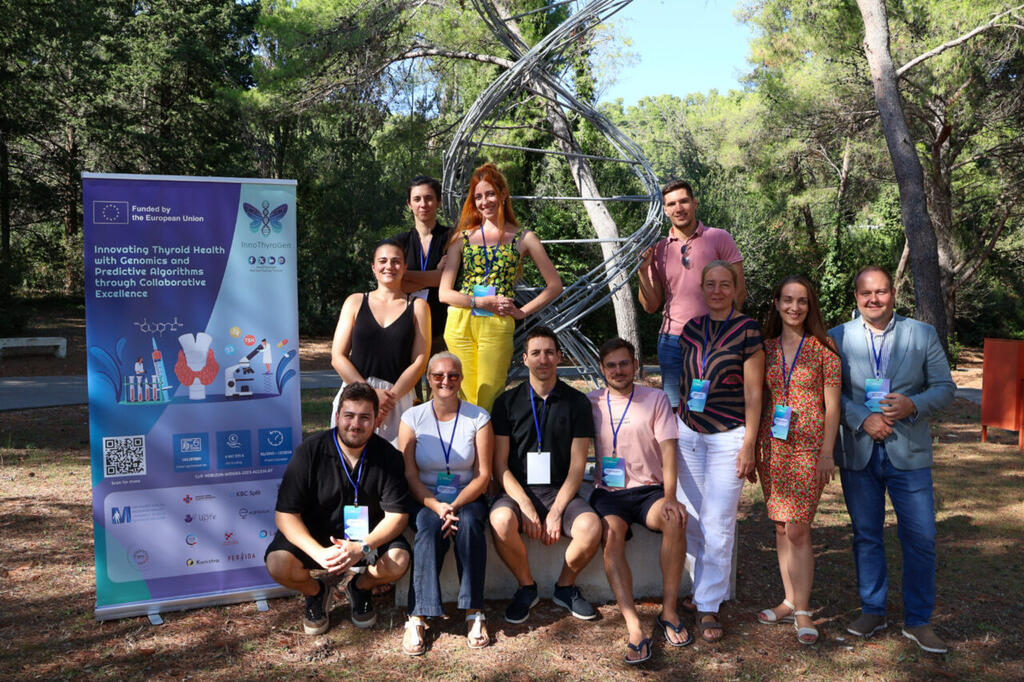
Five days of coding, genomics, and pharmacogenomics brought together 36 young researchers in Split for the first InnoThyroGen Summer School of Bioinformatics.
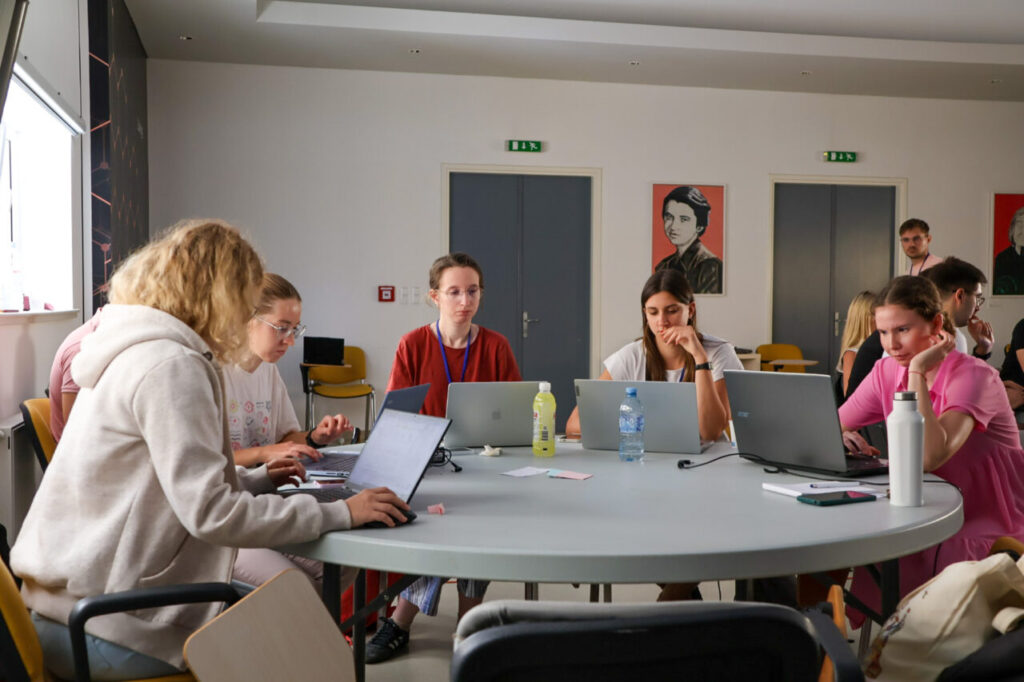
The first InnoThyroGen Summer School of Bioinformatics took place from September 1–5, 2025, at the Mediterranean Institute for Life Sciences (MedILS) in Split. As part of the Horizon Europe project InnoThyroGen, the event brought together 36 participants, including students, PhD candidates, and postdoctoral researchers.
Organizers designed the program to combine theory with practice. This approach gave participants the chance to strengthen their computational skills and immediately apply them to genomics and pharmacogenomics research.
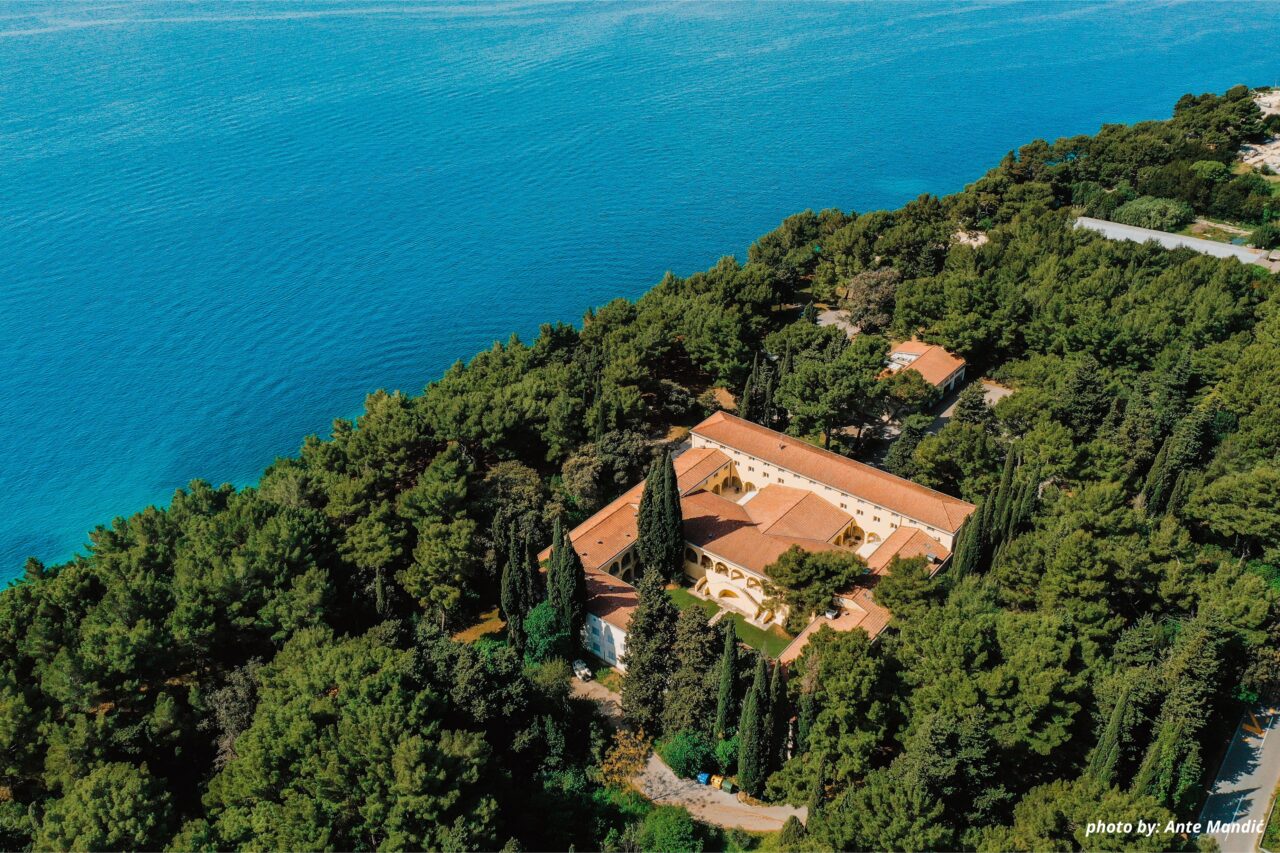

Introduction: InnoThyroGen Summer School of Bioinformatics 2025 in Split
The Summer School was the first of three planned events within the InnoThyroGen project. Over five days, participants gained insights into bioinformatics tools, coding, genomics workflows, and pharmacogenomics applications.

Day 1: Python Basics and AI in Coding
The Summer School opened with welcome addresses by Andrea Gelemanović (MedILS UNIST & MEFST) and Nikola Kotur (IMGGE). After that, Nikolina Pleić (MEFST) introduced the InnoThyroGen project.
Later, Novak Martinović (Persida) presented the basics of Python programming, and Petar Kekez (Sharp Agency) explained how artificial intelligence supports coding. The first day remained more theoretical, which helped participants gradually prepare for the intensive days ahead.



Day 2: Hands-On Python, NLP and Large Language Models
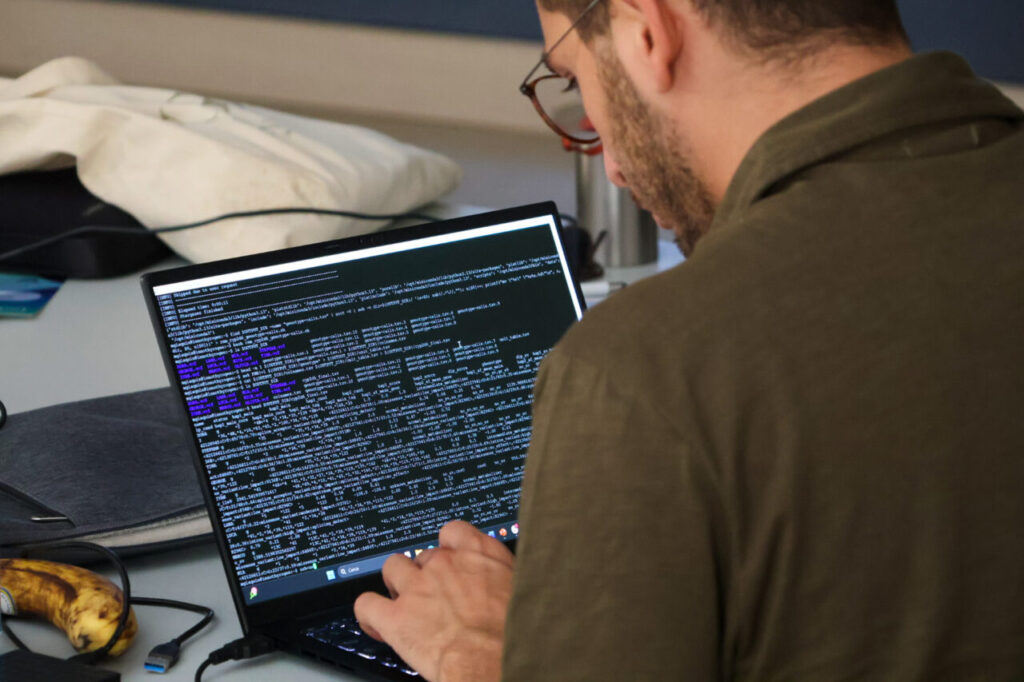
On the second day, participants spent the entire program on Python coding. Nevena Vučinić and Novak Martinović (Persida) guided them through hands-on exercises.
In addition, sessions explored Natural Language Processing (NLP) and Large Language Models (LLM) for biomedical research. The intensive coding was demanding. However, it also became one of the highlights of the Summer School.


Day 3: Genomics Data Analysis and Linux Training
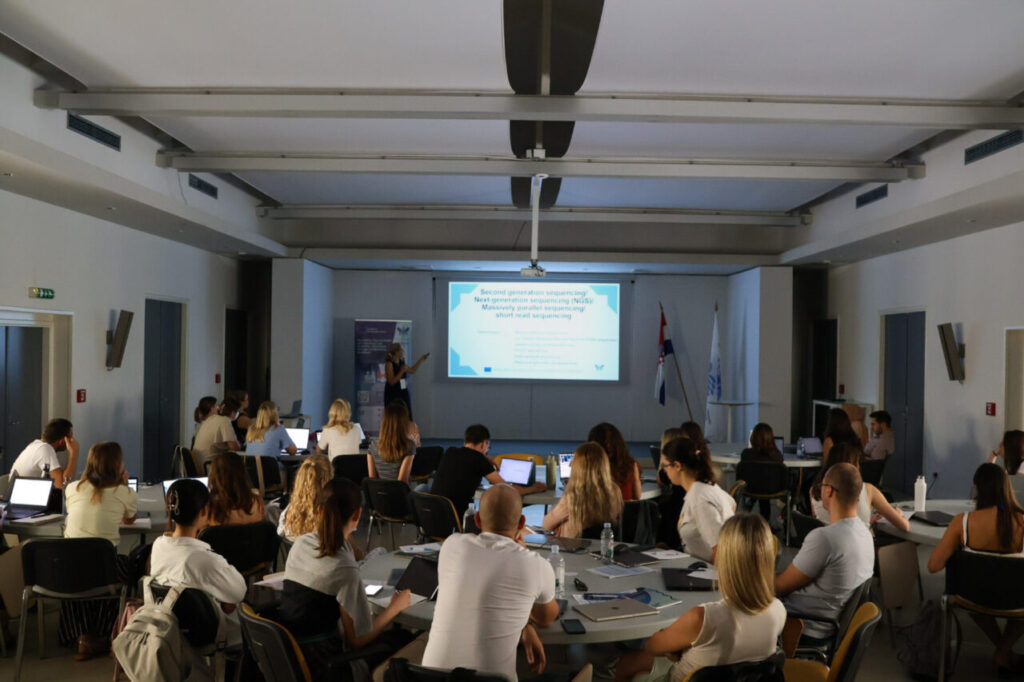
Wednesday shifted the focus to genomics workflows. Branka Zukić (IMGGE) presented the next-generation sequencing workflow. After her session, Nevena Vučinić (Persida) introduced genomics data analysis with NextFlow.
Moreover, Biljana Stanković (IMGGE) delivered a popular lecture on Linux basics. In the afternoon, Đorđe Pavlović (IMGGE) led hands-on sessions on mapping raw reads and variant calling, which participants found extremely useful.



Day 4: Pharmacogenomics Tools and Practical Applications
Thursday focused on pharmacogenomics (PGx). Branislava Gemović (C4IR) explained genomic repositories, followed by Nikola Kotur (IMGGE) with PGx databases and bioinformatic tools. Later, Marina Jelovac (IMGGE) presented population pharmacogenomics.
In the afternoon, Marina Jelovac also demonstrated the Stargazer tool. Afterward, Biljana Stanković and Nikola Kotur introduced the PharmCAT tool. Both sessions stood out for their practical applications.



Day 5: Variant Annotation, HLA Typing, and Future Skills
The final day began with Iva Sabolić (Labena), who presented variant annotation. Next, Mihajlo Stašuk (IMGGE) joined virtually to speak about bioinformatic tools for HLA typing. Finally, Andrea Gelemanović (MedILS UNIST & MEFST) highlighted opportunities for advancing bioinformatic skills.
Closing remarks by Andrea Gelemanović and Nikola Kotur brought the program to an end. Their messages underlined the importance of collaboration and continued learning.
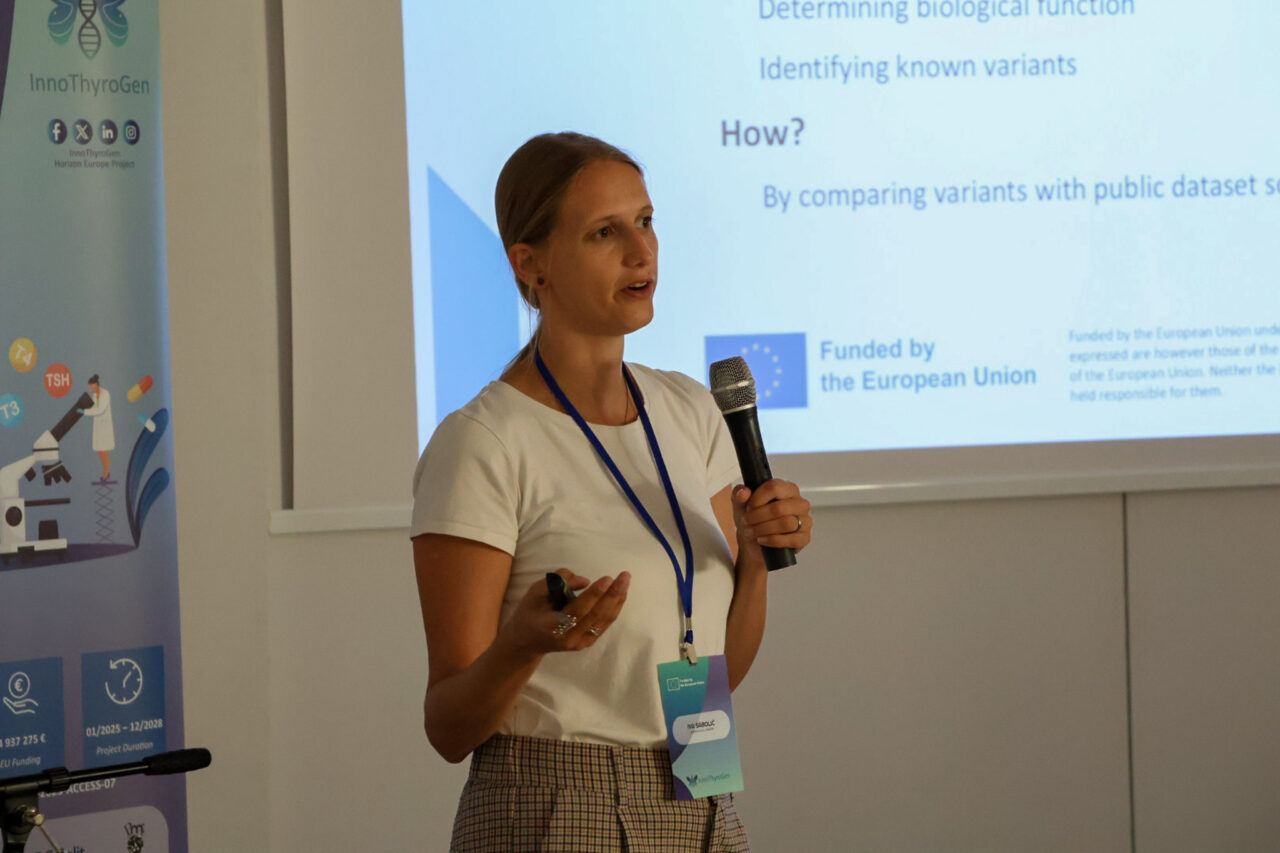
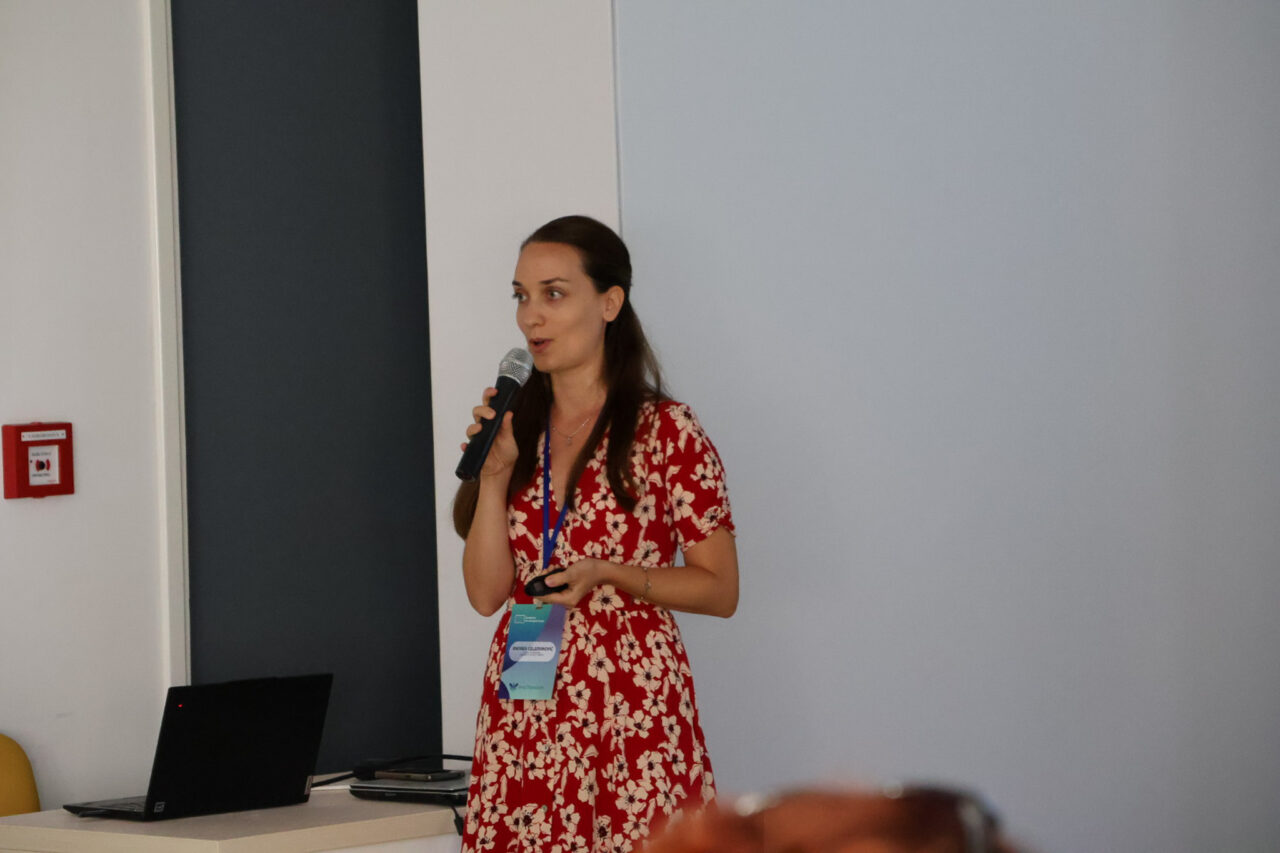



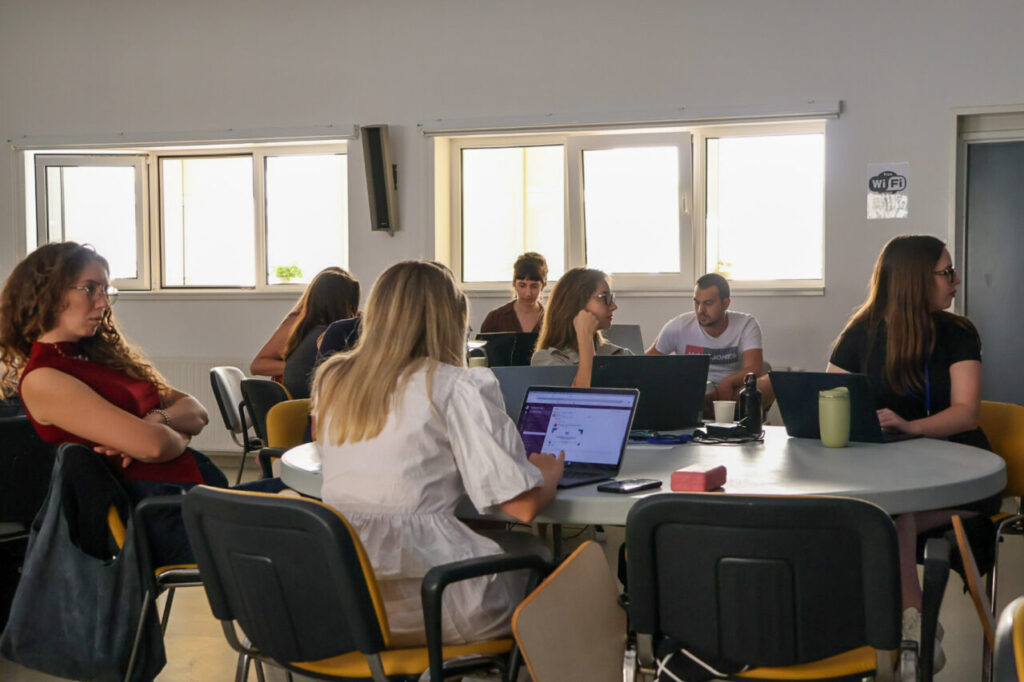
Teamwork and Networking at the Summer School
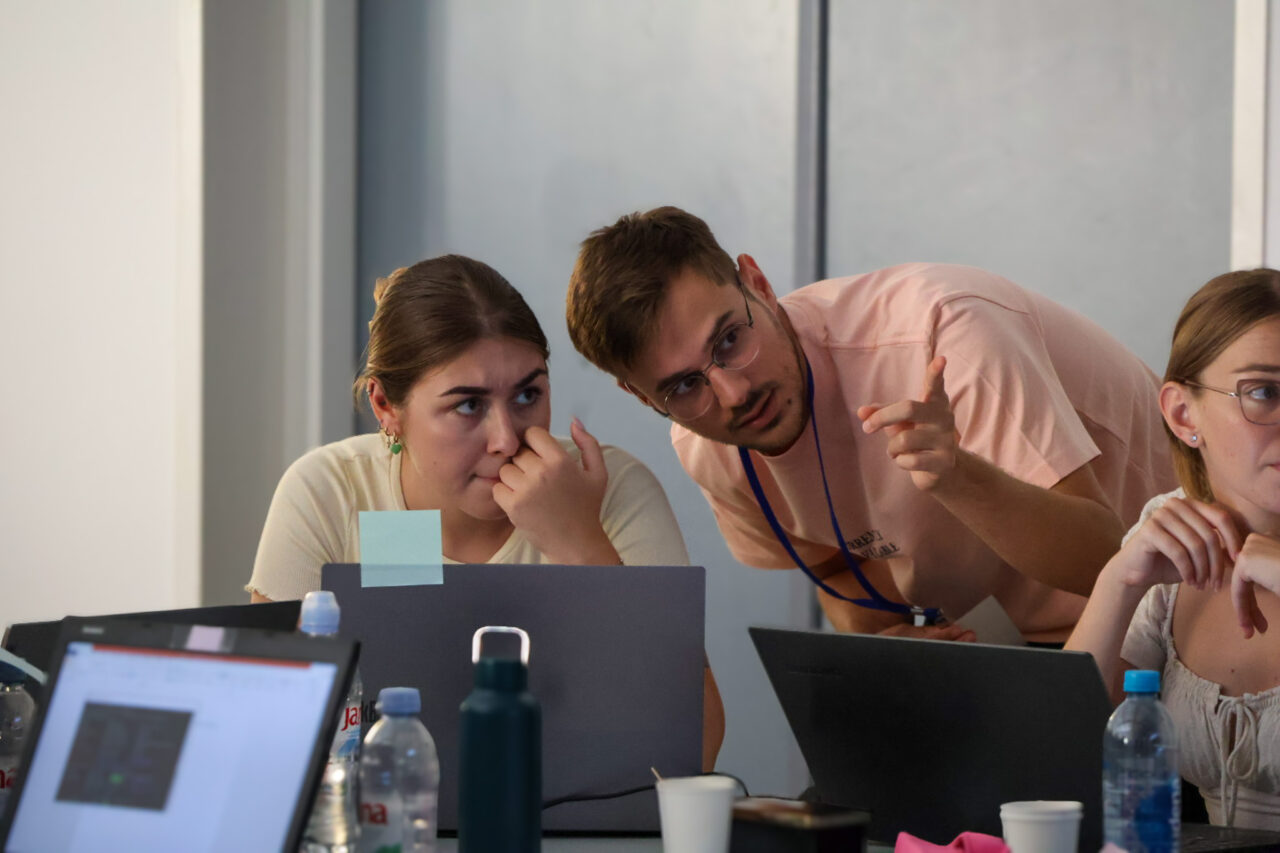
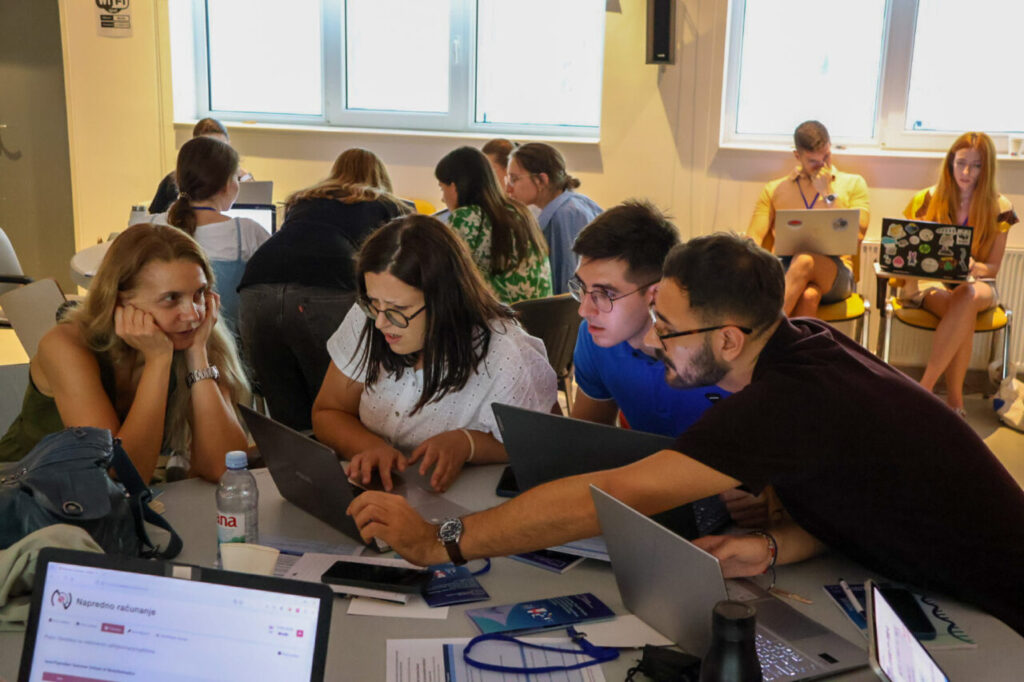
Throughout the week, participants supported one another during exercises. As a result, teamwork became a defining feature of the event.
It was clear that while theory provided a foundation, participants preferred the hands-on sessions. These activities allowed them to apply knowledge directly, which kept engagement levels high.
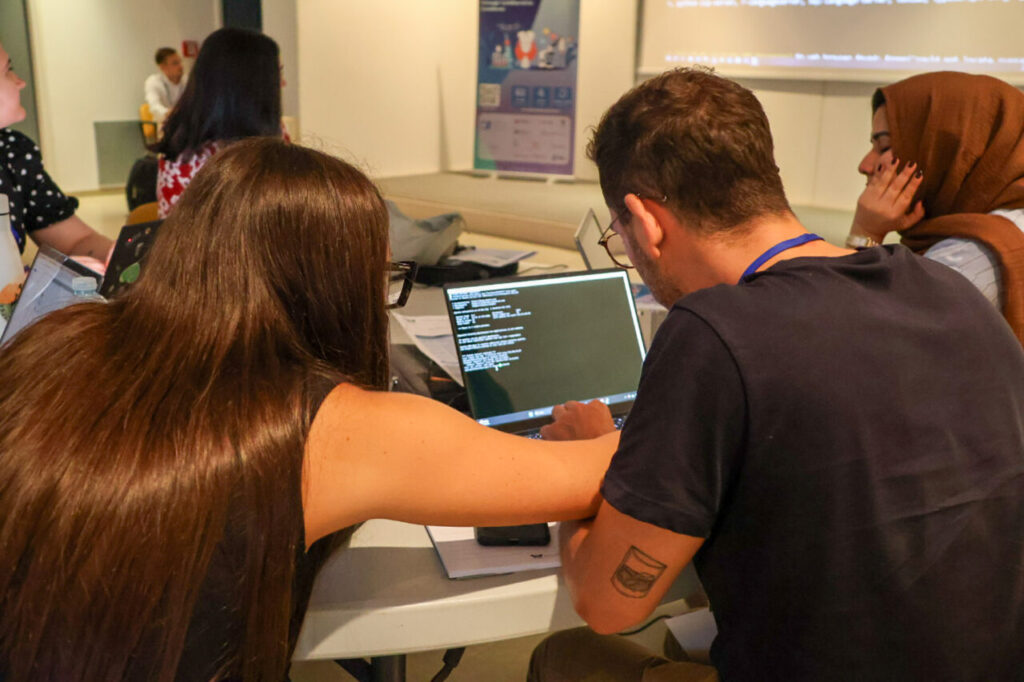



Looking Ahead: More Summer Schools in Bioinformatics and Pharmacogenomics
The success of the first InnoThyroGen Summer School confirmed the value of combining computational training with biomedical research.
Organizers thank all participants for their dedication and all lecturers for their contributions. This was the first of three planned Summer Schools within the InnoThyroGen project.
The next editions will build on this momentum and offer new opportunities for young researchers to strengthen their skills in bioinformatics and pharmacogenomics.
📢 Stay tuned for updates, follow our channels, and don’t miss the chance to join the next InnoThyroGen Summer School!



























